OrthoFinder Tutorials Phylogenetic orthology inference for comparative genomics
2. Running an example OrthoFinder analysis
In the first tutorial you downloaded OrthoFinder and checked you could run it on the Example Dataset. Now you’re ready to run our own analysis!
Plan for this tutorial
In this tutorial we’re going to download the proteomes for a set of species we want to analyse, tidy the files up a little and run OrthoFinder on those species. In the next tutorial we’ll explore the results.
Downloading proteomes for our species
We’re going to perform a phylogenomic analysis across a set of model species: mouse, human, frog, zebrafish, Japanese puffer (Takifugu rubripes) and fruit fly (Drosophila melanogaster). If you’ve returned to this tutorial to guide you through your own analysis you may want check out the advice on species selection in the post, OrthoFinder best practices.
-
Create a folder called “proteomes”. It’s best if there aren’t any spaces in the full path for this folder (e.g. “/home/david/orthofinder_tutorial/proteomes/” and not “/home/david/orthofinder tutorial/proteomes/”).
-
Go to https://www.ensembl.org/, this is generally the first place to look for proteomes. Click on “Human” under “Favourite genomes”. (If you’re downloading data from other websites you might find this post useful: Getting OrthoFinder input data)
-
OrthoFinder requires as input the amino acid sequences for all the protein coding genes in your species of interest. The sequences for each species should be in a separate file with filename extension “.fa”, “.faa”, “.fasta”, “.fas” or “.pep”. When a genome of a species is sequenced and made available, two major steps are performed, assembly and annotation. Assembly is the piecing together of the individual reads into the genome sequence. Annotation is the identification of features of interest in the genome assembly, such as protein coding genes. Therefore, the files we need will often be in a section called ‘annotation’. On Ensembl, on the right hand side, under “Gene annotation” click “Download FASTA”.
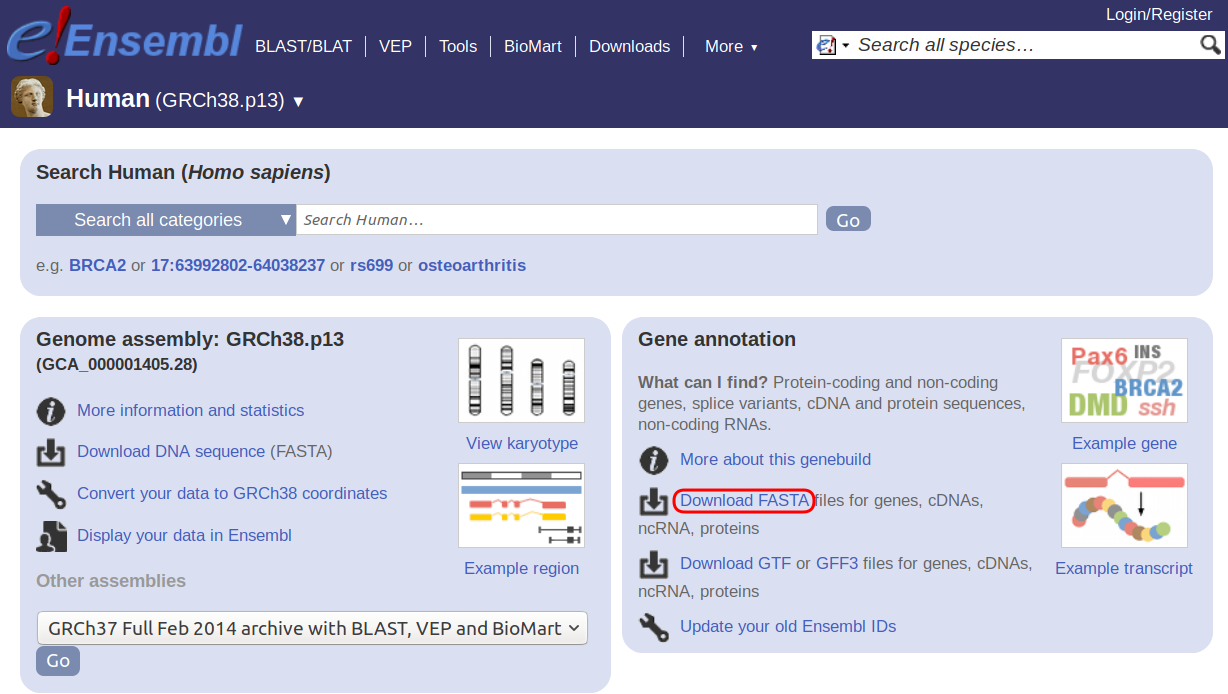
-
Click on the “pep” folder (which contains the peptide sequences) and then download the file “Homo_sapiens.GRCh38.pep.all.fa.gz” to the folder you created.
-
Go back to the Ensembl main page and repeat for Mouse, Zebrafish, Tropical clawed frog (Xenopus tropicalis, under ‘Amphibians’), Takifugu rubripes and Drosophila melanogaster. If there is a choice of files, choose the ‘.pep.all.fa.gz’ file.
-
Open a terminal and navigate to the directory that you downloaded the files to. The files are all compressed (they end in ‘.gz’), decompress them all using the command
gunzip *.gz. - The files from Ensembl will contain many transcripts per gene. If we ran OrthoFinder on these raw files it would take 10x longer than necessary and could lower the accuracy. We’ll use a script provided with OrthoFinder to extract just the longest transcript variant per gene and run OrthoFinder on these files:
for f in *fa ; do python ~/orthofinder_tutorial/OrthoFinder/tools/primary_transcript.py $f ; doneShortening the filename is also a good ideas as it keeps the results tidy as the filenames are used to refer to the species, e.g. I shorten to Homo_sapiens.fa.
- Run OrthoFinder (if you’ve not performed the above steps yourself but would like to run OrthoFinder you can get the prepared files here: primary_transcripts.tar.gz.)
orthofinder -f primary_transcripts/ - That’s it! It will take from around 20 mins to a few hours to run, depending on how many cores you have and the speed of your machine. Assuming everything worked fine then the last few lines of OrthoFinder’s output will look something like this:
Results:
/home/emms/orthofinder_tutorial/proteomes/primary_transcripts/OrthoFinder/Results_Nov26/
OrthoFinder assigned 121743 genes (92.9% of total) to 17981 orthogroups. Fifty percent of all genes were in orthogroups with 7 or more genes (G50 was 7) and were contained in the largest 5076 orthogroups (O50 was 5076). There were 5485 orthogroups with all species present and 1755 of these consisted entirely of single-copy genes.
CITATION:
When publishing work that uses OrthoFinder please cite:
Emms D.M. & Kelly S. (2019), Genome Biology 20:238
If you use the species tree in your work then please also cite:
Emms D.M. & Kelly S. (2017), MBE 34(12): 3267-3278
Emms D.M. & Kelly S. (2018), bioRxiv https://doi.org/10.1101/267914
That’s it! The next tutorial is: Exploring OrthoFinder’s results. If your OrthoFinder run hasn’t finished then you can still continue with the next tutorial, it contains a link where you can download an archive of the results.
Written on September 19th, 2019 by David Emms